Image orientation in CamiTK
CamiTK is dedicated to prototyping application in the domain of Computer Assisted Medical Intervention (CAMI), sometimes called Computer Assisted Surgery. The main target are scientists and companies that develop innovating medical devices driven by software. Clinicians can also use CamiTK in order to visualize medical data and/or to apply some image processing. It involves 2D and 3D images and interaction of these images with other components.
Images orientation has 2 purposes:
- slice visualization and
- images registration with other components.
Slice Visualization
In order to allow clinicians to go through 3D images their usual way (using Axial, Coronal and Sagittal slices), we had to find a way of orienting our images in order to visualize them.
Viewing conventions
There are two widely used viewing conventions: 1
- Radiological convention: Images are viewed as though looking upward from the feet of the subject (feet to head). “Right is Left”
- Neurological convention: Images are viewed as though looking from the top of the head downward (head to feet). “Right is right”
The DICOM standard does not specify one of this viewing convention as the one to use. But, as most users were using the radiological convention, we decided to use this one in order to visualize all our images in CamiTK.
DICOM default orientation
DICOM defines a term, which will help choosing an orientation: “Reference Coordinates System” or RCS 2. The RCS is a very intuitive coordinate system of the patient body: X direction is from Right to Left. So if the patient is standing in front of you with the arm raised to the sides, then X direction is from the right hand to the left hand."


The patient is laying on the imager table and the clinician looks from its feet.
Orientation is RAI: Right first, then Anterior first, then Inferior first. We look at the patient from his/her right (his/her right hand is the first to appear on the Coronal slice), then to his/her left, then from his/her front (nose) to his/her back and then from his/her feet to his/her head.
CamiTK convention
CamiTK does not specify a convention anymore. If the loaded image provides the anatomical information, CamiTK will label each axis accordingly (e.g. L/R, I/S, A/P). If no information is provided, it will display X, Y and Z for each axis. Each image has a [Frame Of Reference](Frames and Transformations/) which can be edited to set anatomical labels to the axes.
Before version 6, CamiTK followed the RAI orientation, meaning that all volume images (ImageComponent instances) would be oriented in order to get the image coordinate system in the RAI position, such as showed by this picture:

CamiTK slices visualization
The Axial, Coronal and Sagittal viewers are tagged with patient position (Anterior / Posterior, Superior / Inferior, Right / Left) in order to locate more easily each slice of the image.
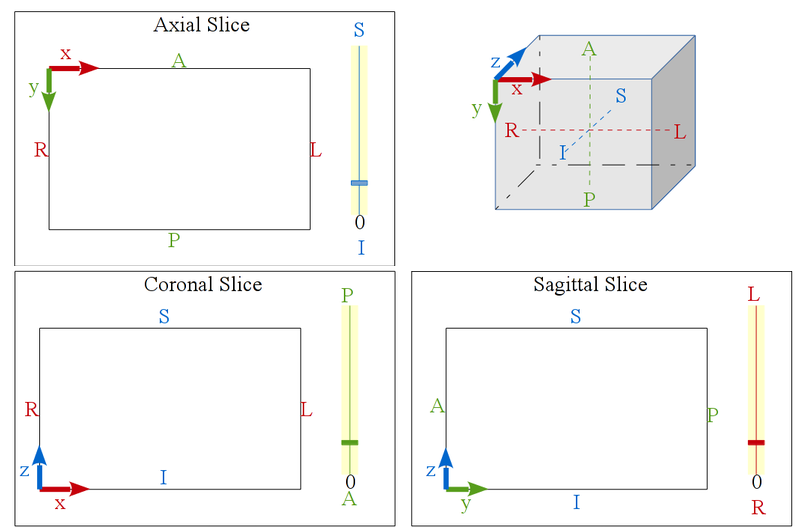 |
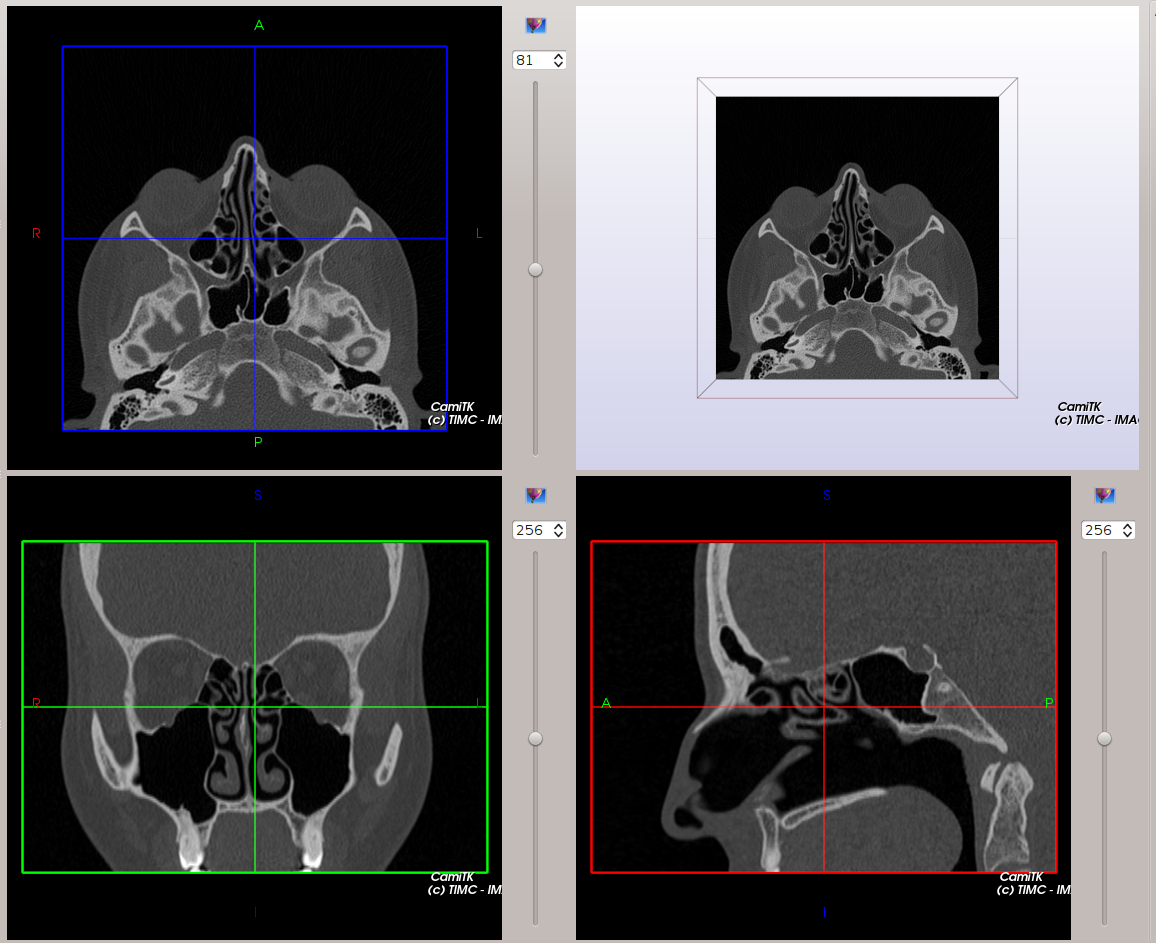 |
Our MedicalImageViewer is inspired from Slicer3.
CamiTK DICOM visualization
DICOM is a file format that can store volumic images, read from a directory containing several files of a serie, belonging to a study. In order to correctly orient the image, the following DICOM tags are taken into account:
- (0020, 0037) Image Orientation Patient: also called “Direct cos angles” this information allow us to deduce the rotation from the image coordinate system (what was the orientation of the patient when the image was acquired) to the RAI coordinate system.
- (0020, 0032) Image Position Patient: gives the position of the first voxel of the image. Allows to deduce the translation from the scanner coordinate system to the patient coordinate system (RAI). By default, the scanner coordinate system is the main frame of the image, its data frame being the untransformed voxel matrix frame.
Technical information
Image orientation is now managed by the Frames and Transformations system: images are loaded ‘as is’ by the relevant Component, and the ImageComponent created while opening the file has two FrameOfReference: the data frame which is directly linked to the voxel matrix, and the main frame, which provides a Frame specified by the file format (e.g. the transformation to a ‘scanner-based’ frame for a CT-scanner or an MRI). The function that creates the Component should set the Anatomical information if it is available, using the FrameOfReference’s AnatomicalInformation object. This allows CamiTK to know which labels should be displayed on slices. For the moment, CamiTK does not change the orientation of the displayed slices even if the expected ‘Axial’ slice is not axial, but the labels will be correct. Please take note that the Frame anatomical information will be saved in a CamiTK workspace, but not in the usual image formats (mha, nii…).
Saving an image
VTK does not support frames. If you try to save both your image and its frame using the header of MHA file, there will be some errors on the data. To avoid this problem, we choose to save only the basic VTK image, without its frame transform.